 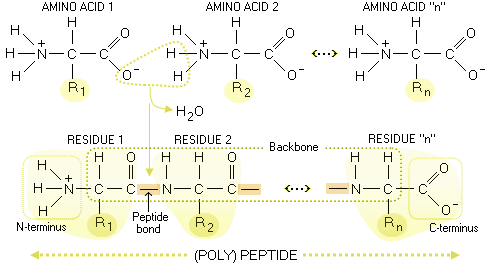
which bind tightly to the negatively charged DNA backbone. This chapter includes the structure, functions and stereochemistry of peptide bonds. One end is positively charged, while the other is negative. Peptide bonds are the basic backbone of the proteins. A deep understanding of chemistry is essential to anyone interested in modern biological sciences or medicine, so I really encourage you to take the time to work though all of the chemistry material. One end contains the telomere, while the other contains the centromere. That may look like a lot of work, but you've probably watched many of the videos already under "Chemistry of life". If you are not familiar with reduction and oxidation states, then I encourage you to start working through the Chemistry material on KhanAcademy: How many moles of carbon are present in a gram of glucose (a "typical" carbohydrate)?ĭo the answers to those two sets of questions help you answer your question? How many moles of carbon are present in a gram of tetradecane - a 14 carbon alkane (a reasonable comparison for the tails of the fatty acids found in food)? How would you expect this to affect a oxidative process like cellular respiration? This is because moving an angle in the middle of the protein can result in large movements at the end of the protein (think of it like putting a small bend in. How does this compare with the oxidation state of the carbon in carbohydrates (the other group of macromolecules that are often used to store energy)? The N-terminus is on the left and the C-terminus is on the right for the very short polypeptide shown above. The other end, which has a free carboxyl group, is known as the carboxyl terminus (or C-terminus). What is the oxidation state of the carbons in the fatty acid tail? Surprisingly, the frozen structures from ultra-high-resolution protein crystallography reveal a prevalent, but subtle, mode of local backbone motion coupled. At one end, the polypeptide has a free amino group, and this end is called the amino terminus (or N-terminus). Therefore, I'm going to ask you some questions in response to help you figure out (some of) the answers yourself. Together, these results suggest a mechanism for fast protein structure refinement and folding using external energy functions.This is a good question, but one that I think you have enough information to answer on your own. Finally, we perform sequence design, relaxation, and ab initio folding of a subset of generated structures, and show that in some cases we can recover the generated folds after forward-folding. We show that interpolations in the latent space of the generator correspond to smooth deformations of the output backbones, and that test set structures not seen by the generator during training exist in its image. These classifications may differ in the number of structural classes, the length of peptides, or class attribution criteria. We represent protein structures by pairwise distances between all backbone atoms, and present a method for directly recovering and refining the corresponding backbone coordinates in a differentiable manner. Motivation Local protein structure is usually described via classifying each peptide to a unique class from a set of pre-defined structures. In this study, we train Generative Adversarial Networks (GANs) to generate fixed-length full-atom protein backbones, with the goal of sampling from the distribution of realistic 3-D backbone fragments. Abstract: The fast generation and refinement of protein backbones would constitute a major advancement to current methodology for the design and development of de novo proteins. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
